Virtual Screening with AI Agents
Deploying our agentic platform to accelerate functional workflows
Axon is an interactive AI platform that plans, executes, and analyzes computational chemistry tasks via natural language. Validating Axon’s utility in drug discovery necessitates examining its function at scale. As a first case study, we walked through a virtual screening example that explores how Axon can be used to filter thousands to millions of molecules down to a few promising drug candidates in a matter of hours. Here, we emphasize Axon’s role as an orchestrator and focus on accelerating rigorous deployment of existing tools and benchmarks, not on the quality of the underlying tools themselves.
DUD-E Docking Benchmark
The chemical space for drug discovery is enormous, and experimental screening of millions of compounds is prohibitively time- and cost-intensive. Virtual screening is increasingly becoming an essential part of integrated drug discovery campaigns, where it is used to identify promising molecules that proceed to more expensive experimental validation stages.
Structure-based virtual screening relies heavily on molecular docking, a computational method that predicts how well a small molecule ligand (drug candidate) binds to a target protein. This typically involves sampling different positions in which the ligand can sit in the protein binding pocket and scoring the energetics of each pose.
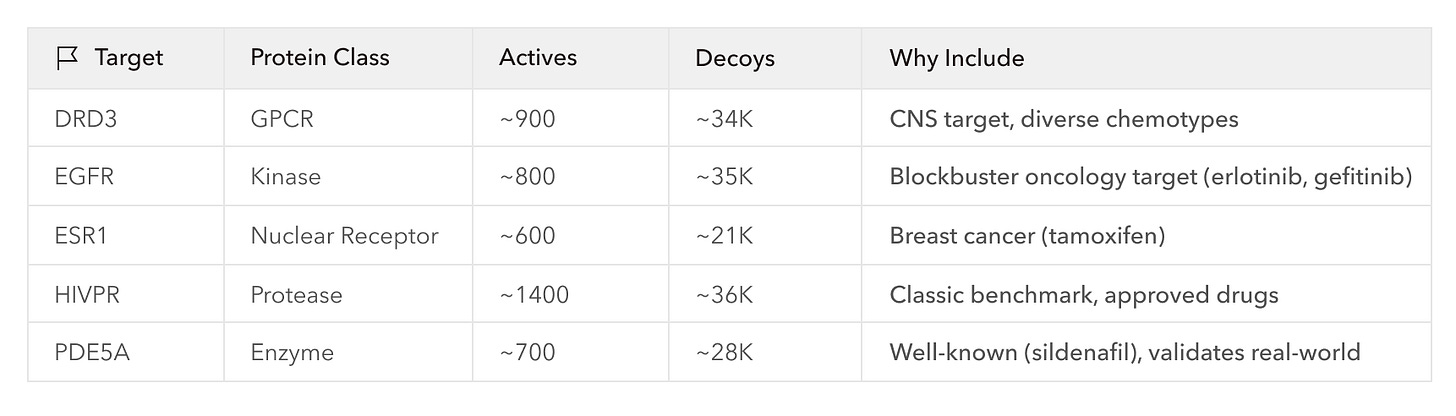
There are many different approaches to molecular docking, and benchmarks have been developed to compare them. One of the most commonly used benchmarks is the DUD-E (Directory of Useful Decoys, Enhanced) dataset comprising 102 diverse protein targets. Each protein is paired with hundreds of active ligands (224 on average), and to emulate the challenge of drug discovery, DUD-E provides 50 property-matched but inactive “decoy” ligands for every active ligand. Docking programs are tasked to rank the blindly mixed actives and decoys by their binding activity, and are evaluated on enrichment factor (EF), a measure of how concentrated the actives are at the top of their ranked lists, and AUC-ROC, which measures true positives against false positives through all possible score cutoffs.
The Gap
Beyond the time demands and computational cost of screening millions of molecules, the setup and analysis of new screening campaigns are among the most significant bottlenecks in virtual screening. For example, while docking tasks themselves may seem straightforward, they actually require many steps of preparatory work including downloading large volumes of data, cleaning and formatting the data, configuring HPC infrastructure, and identifying appropriate screening parameters. And as any computational scientist knows, every step comes with potential for a long tail of ancillary work in troubleshooting and de-bugging, especially when implementing at scale.
While agentic systems have not yet been systematically evaluated for computational benchmark setup, there is significant precedent that suggests substantial acceleration potential, including successful deployment of LLM agents in adjacent scientific domains and demonstrated LLM capability in closely aligned software-engineering activities: file format conversion, dependency resolution, parameter configuration, error debugging, etc.
Axon’s Role
This is where Axon comes in. As an agentic research environment, Axon can understand your goals, systematically compile relevant information, reason through errors autonomously, and execute tasks at scale. This frees up users to focus on understanding the science: asking meaningful questions, and setting direction to keep research moving forward. Here, Axon enabled us to strategize around, set up, and run a subset of DUD-E end-to-end in hours.
Importantly, Axon is an interactive program—the scientist is always in the loop. If we had been very familiar with DUD-E and knew exactly what needed to be done, we likely could have provided a detailed one-shot prompt instructing Axon to execute the whole process in a single go. Instead, however, we proceeded by collaboratively making a plan with Axon, deciding to run a two-step virtual screening workflow on a subset of DUD-E covering ~155K docking calculations across 5 diverse, pharma-relevant targets; just enough to demonstrate correct and scalable execution of the benchmark. The workflow involved two established docking platforms: a first-pass ranking of the DUD-E ligands with AutoDock-GPU, then further enrichment of the top 1% of the ranked ligands with GNINA, a more expensive, ML-enhanced method.
Ultimately, it took a bit of tuning and inspection, but Axon was able to set up and run the docking workflow successfully. The TL;DR is that Axon measured AutoDock-GPU AUROC scores comparable to published literature values for each target and demonstrated considerable enrichment of the top ligands with GNINA, clearly accelerating the process by handling details and infrastructure related to data processing and tool use. However, Axon also encountered a few hiccups that made the value of having expert input clear, and highlighted a distinct roadmap to improvement including teaching Axon with usage and feedback and improving tool integrations.
The Details
We actually ran the DUD-E workflow end-to-end twice. The first time, we operated Axon with very little oversight over it’s activities and provided only high-level instructions (i.e. goals and workflow overview). We loosely sanity-checked Axon’s plans and outputs, but let Axon handle all of the strategy, data retrieval, tool selection, and compute allocation. Axon ran through the workflow quite smoothly, seemingly handling minor snags such as retrieving files directly from RCSB PDB when it found that the DRD3 protein target file was inaccessible from the DUD-E archive. Looking at the results, however, the performance was clearly poor: the AUC scores for AutoDock-GPU were low (AUC=0.5 is equivalent to randomly ranking the ligands) or even inversely correlated with our ranking (AUC<0.5).
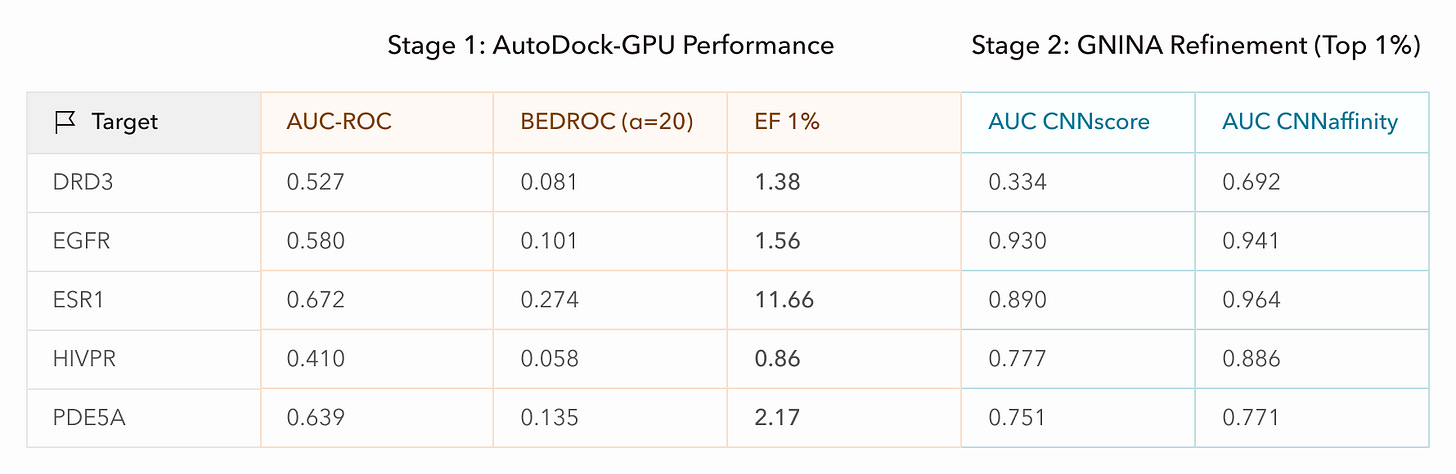
After probing Axon for feedback and performing some manual inspection, we identified a few issues:
The DUD-E website experienced intermittent connectivity issues that hindered Axon from retrieving data smoothly. As a result, Axon turned to PDB and retrieved a slightly different version of DRD3that had a mis-matched docking center.
Axon only found SMILES strings for the decoy ligands of a few targets, so it generated its own 3D conformations from these, resulting in inconsistent preparations. This is likely attributable to the connectivity issues preventing Axon from being able to search the archive thoroughly.
Axon used sub-optimal filetypes and tools for ligand pre-processing based on what was pre-installed (MOL2 filetypes, OpenBabel) despite having the ability to ‘pip install’ other packages. The ideal tool chain for AutoDock-GPU utilizes SDF file formatting, which DUD-E also provides, and MGLTools, which is developed by the creators of AutoDock-GPU and better ensures compatible atom type mapping, partial charge calculation, etc.
Notably, these are missteps a domain expert likely would have identified immediately, or would have circumvented altogether by checking the input files against available files and indicating their preferred tools.
Starting again with a clean session, we pointed Axon to specific files from DUD-E, including SDF-formatted ligand files, and requested that it use MGLTools for receptor and ligand pre-processing, which includes file conversion (AutoDock-GPU uses PDBQT format) and Gasteiger partial charge calculations. We also requested that Axon parallelize docking for each receptor over 32 GPUs, bringing wall-clock time down to half an hour for the run. We let Axon use default parameters for everything else. Immediately, AUC-ROC scores were significantly improved and were on-par with published literature values for these targets, demonstrating correct deployment of AutoDock-GPU.
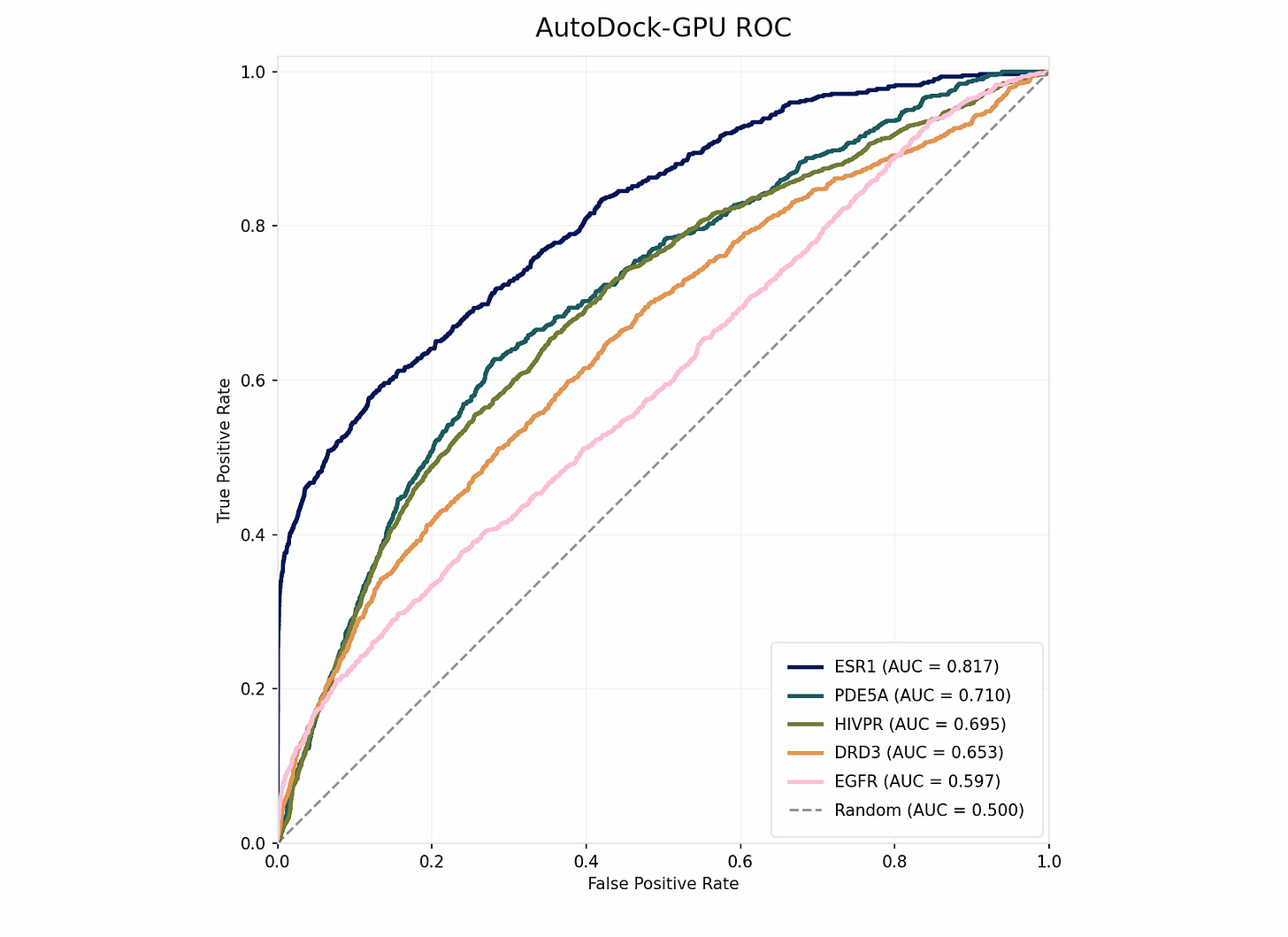
Next, since GNINA previously demonstrated reasonable enrichment without supervision or intervention, we prompted Axon to perform the same GNINA refinement on the top 1% of ligands from AutoDock-GPU using default parameters and parallelized processing. With better initial screening, GNINA also achieved better final screening via both of its outputs, CNNscore (classification-style probability) and CNNaffinity (regression prediction of binding affinity). Targets with weaker-to-moderate enrichment may not have had good representation in the GNINA training data, or may be better suited to other docking methods due to target-specific qualities such as high flexibility.
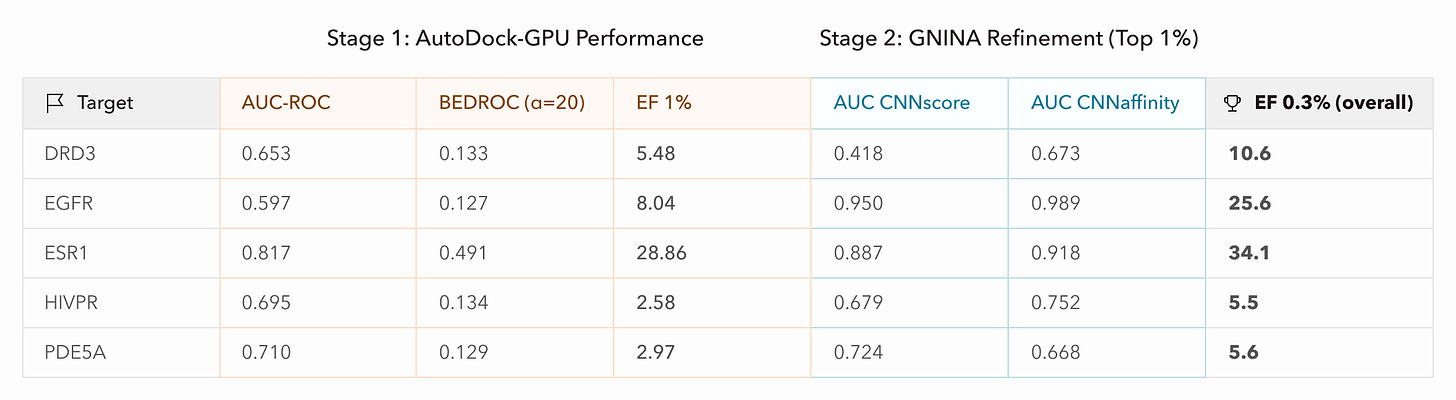
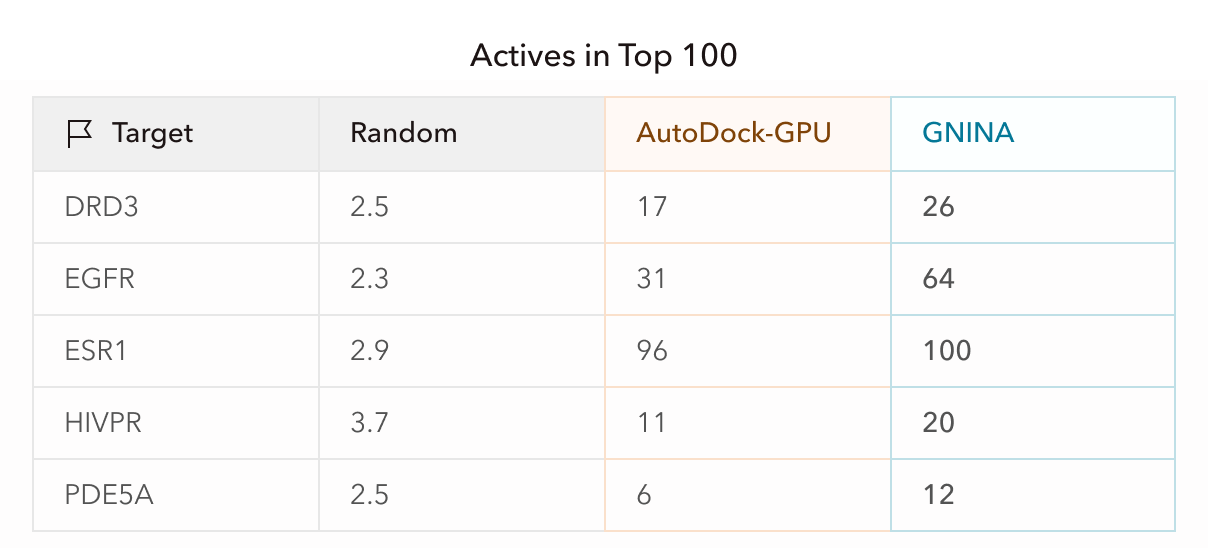
What these results mean practically is that if a small biotech company screened drug candidates and proceeded to test the top 100 molecules experimentally, equivalent to about the top 0.3% of the initial ligand libraries, this screening pipeline would have increased active hits by 5-35x per target. For ESR1, in fact, the top 100 candidates were all active, and GNINA refinement approximately doubled the hit rate for the other targets. This enrichment directly impacts a company’s bottom line and can be notably accelerated with Axon.
Reflections
In this workflow, we were impressed with Axon’s ability to understand scientific goals, find documentation, plan, and execute workflows at production volume. Axon deftly dealt with procedural implementation details such converting between filetypes and piping data around for parallel processing. This in itself, I’m confident, can save computational scientists a lot of time and headache.
Left to its own devices, however, Axon could not “one-shot” the workflow with only a high-level prompt and instead required some oversight of its decisions and body of work. As such, debugging this workflow highlighted the importance of the human-agent interface for scientific collaboration. In Axon, conversation history and data inputs and outputs are all easily accessible, and Axon can be continually probed for what it did and how it made decisions. Iterating on experimental design is a normal part of scientific research—but especially with the expertise of a scientist familiar with domain tooling and data, Axon’s operational autonomy and information provenance made it much faster and easier to drive continual progression.
The hiccups Axon encountered reflect Axon’s nascent stage and outline a clear roadmap for improvement. With feedback from use, and with our continued work on tool integration, we expect that Axon will learn to streamline workflow set-ups more and more rigorously. It might, for instance, adapt heuristics that indicate which file formats are optimally compatible with each other or with specific tools. It’s my belief that in research, expert input and guidance will always be valuable. But the more we can automate implementation mechanics, the more we can free up experts to focus on the scientific work itself: asking hard questions, dreaming big, thinking critically about what comes next.
Try for yourself!
Hopefully it’s clear how Axon can accelerate virtual screening and similar computational processes at scale; notably, by automating tool and workflow setup, as well as by handling resource management and parallelization. If these sound interesting or helpful to you, you can try your own workflows on Axon here!
We're super excited about Axon and have way more to say about it than fits in a blog post. If you have questions or comments or just want to talk with us, shoot me a message at sabrina@mirrorphysics.com.